Example file[edit]
You can download it here -> OptKnock_toy.zip
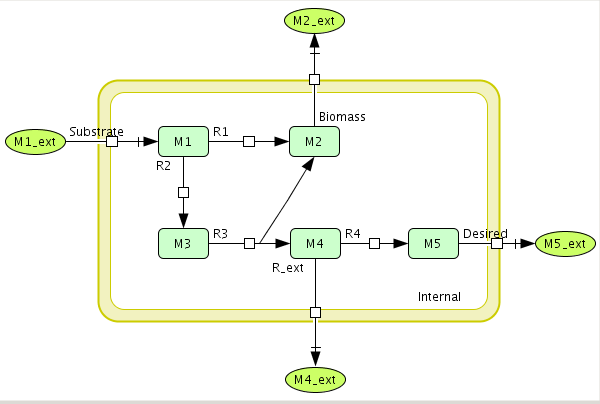
An illustrative image of this model can be found below.
The simple network is constituted of seven reactions. The reactions: Desired, Biomass and substrate represents the production of the biological coumpound of interest, the formation of biomass and the uptake of metabolite M1 from surrounding media, respectively.
The coefficient of reaction R2 (M1->M3) are such that two molecules of M1 create one molecule of M3, while the stoichiometric coefficients of all other reactions reflect a one-to-one relationship between molecule quantities.
Brief tutorial[edit]
This brief tutorial will help you to begin working with OptKnock4OptFlux.
- To start, you will need load a model in OptFlux(see how to here). You must pay attention to select a correct biomass flux.
Run OptKnock[edit]
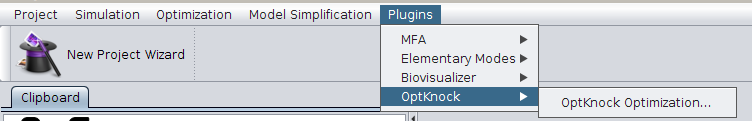
- 1 - Start by accessing the menu Plugins->OptKnock->OptKnock Optimization.
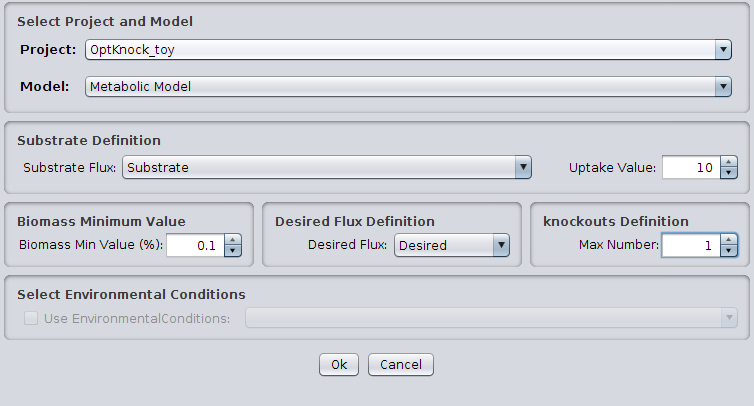
- 2 - The following dialog will be displayed.
1. Select Project and Model
You can select the model/project to work.
2. Substrate Definition
You can select the substrate flux and its uptake value.
3. Biomass Minimum Value
You can define the minimum value for the biomass (in %).
4. Desired Flux Definition
You can select the reaction representing the production of the desired compound.
5. Knockouts Definition
You can define de maximum numbers of knockouts.
6. Selected Environmental Conditions
You can define the environmental conditions if you have created any.
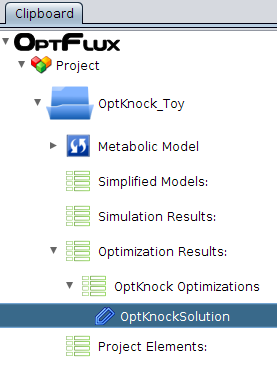
- 3 - After execution, a new datatype will be placed in the clipboard, under the Optimizations Results -> OptKnock Optimizations.
Pressing it will launch three viewers.
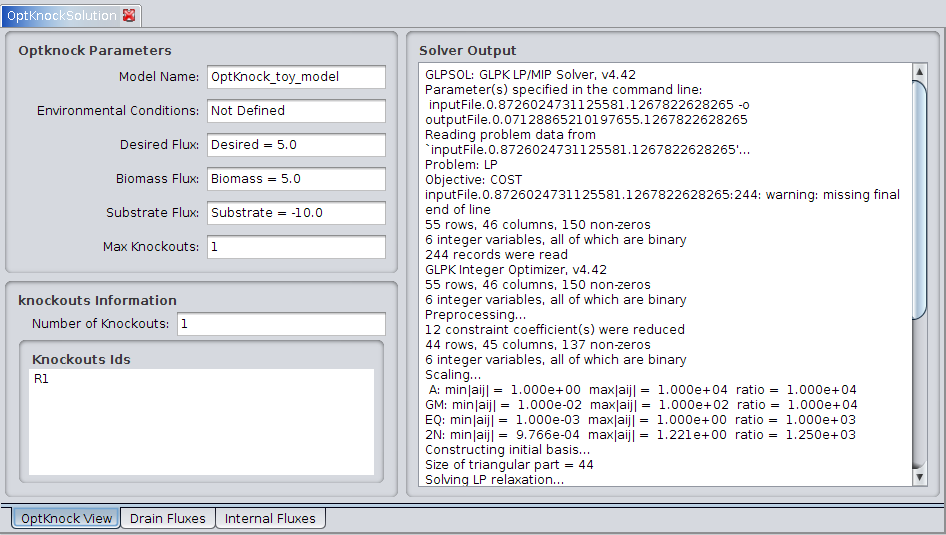
1. OptKnockView
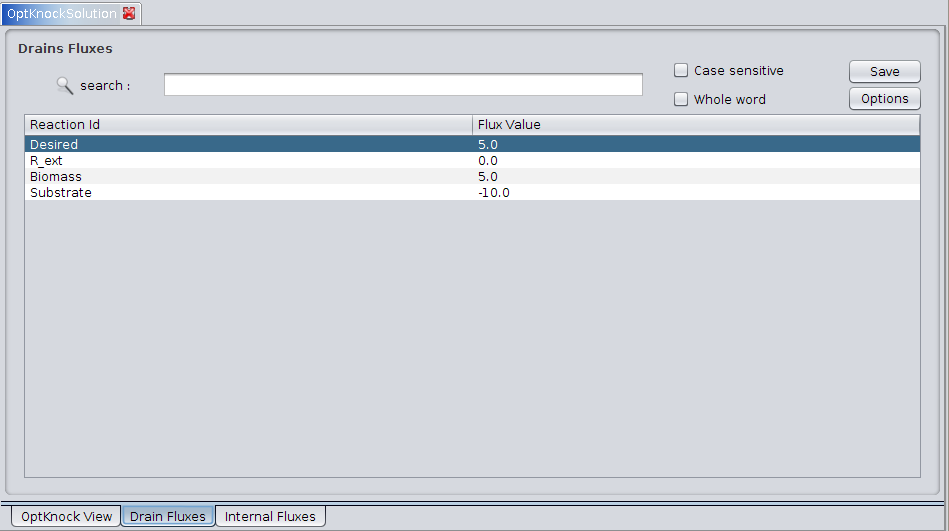
2. Drain Fluxes
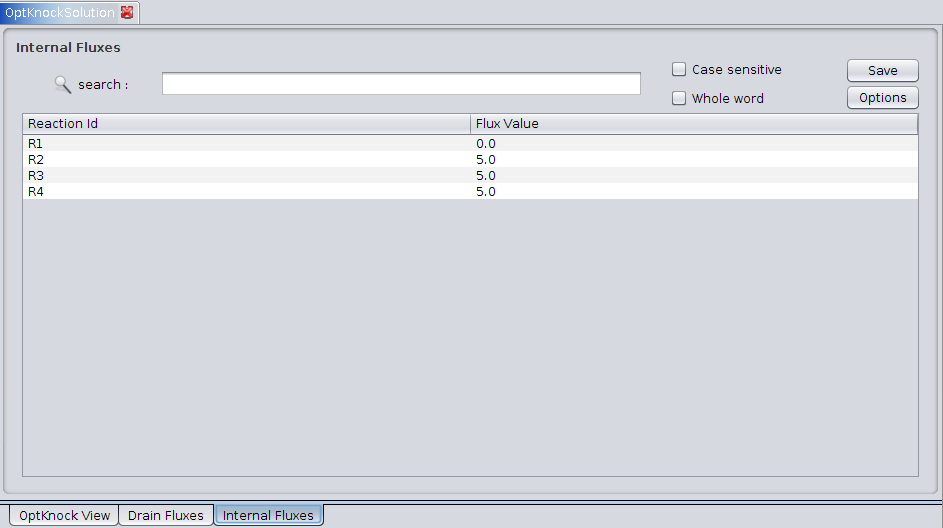
3. Internal Fluxes