Contents
How to create a network[edit]
- 1 - Go to Plugins → TNA4 → New network.
- 2 - Select the project which the network will be associated with.
- 3 - Select the type of network that you which to create, there are three possible kinds of networks: (1) Reaction-Metabolite Networks, (2) Metabolite only Networks and (3) Reaction only Networks.
- 4 - Give a name to your network.
- 5 - Select if drain reactions and external metabolites should be included in the network.
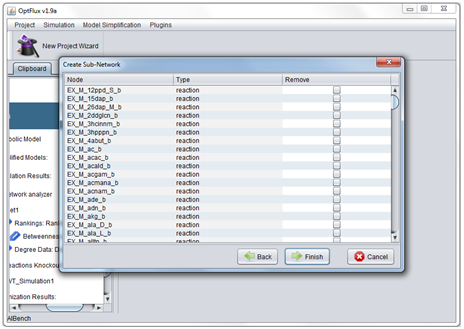
- 6 - If necessary some of TNA4OptFlux's filters during the creation of the network.
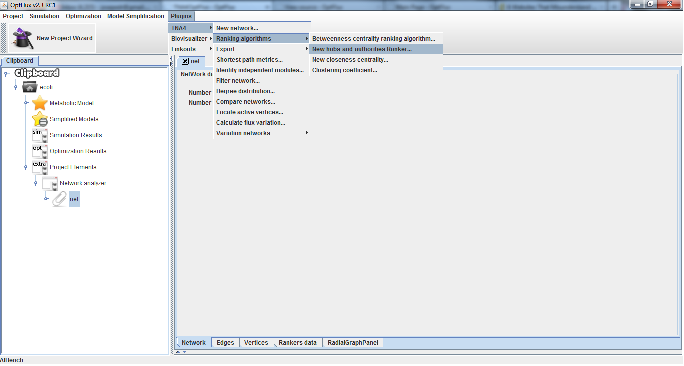
Using ranking algorithms[edit]
- 1 - Go to Plugins →TNA4 → Ranking algorithms, then chose the desired algorithm from the submenu.
- 2 - Results can then be saved to a ASCII file or visualized directly with the application.
Exporting networks[edit]
- 1 - Go to Plugins →TNA4 → Export.
- 2 - Select the desired format.
Shortest path metrics[edit]
- 1 - Go to Plugins → TNA4 → Get Shortest path metrics.
- 2 - Select the network to analyse.
- 3 - After the shortest path metrics are calculated there are four views that can be used to analyze the data.
The four shortest path views and their functions are:
- 1 - Shortest path view - this view shows a few global network metrics based in the shortest path.
- 2 - Shortest path - can be used to calculate the shortest path between a pair of vertices, by default BFS is used if the option "Get all shortest paths" is used SBBFS is used instead.
- 3 - Shortest path vertex data - this view is used to determine the shortest path between two nodes of the network.
- 4 - Shortest path Histogram - this view shows a table containing the number of Shortest paths in the network which have a certain length.
Independent modules[edit]
Occasionally networks contain independent modules, these are sections of the network which are isolated from the rest of the net. TNA4OptFlux can be used to find these subnetwork, and if necessary it can convert them into independent networks so that the user can analyze them apart from the rest of the large network. To indentify the independent modules simply go to Plugins → TNA4 →indentify the independent modules. To convert a independent module into a network simply select that option from the view of the desired module after indentifying it.
Filters[edit]
Filters can be used to create subnetworks though the removal of elements from an original network based in user defined criteria.
To use filters in TNA4OptFlux go to Go to Plugins →TNA4 → Filter Network and use the wizard to define the filters which should be applied to the network.
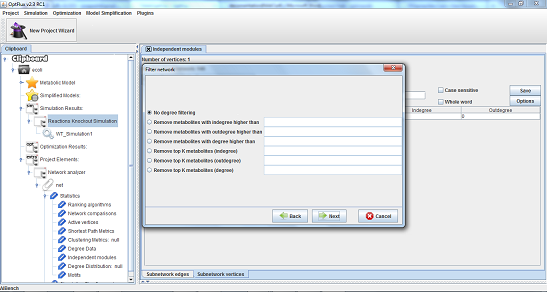
There five filtering options in:
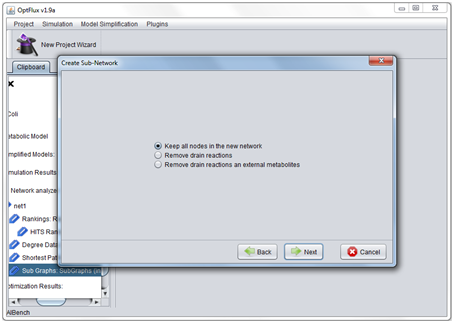
- 1 - The removal of drain reactions and external metabolites.
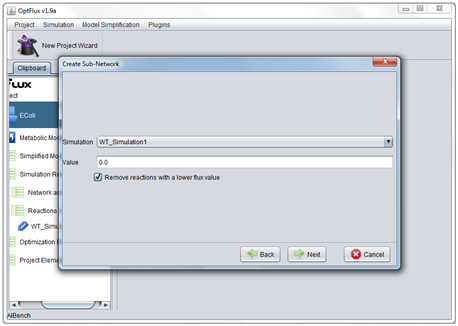
- 2 - If the project has any associated simulations it is possible to use simulation filters which remove from the network all reactions whose simulated flux is bellow a defined threshold.
- 3 - Degree filtering is a option which can be used to remove vertices based in their degree value.
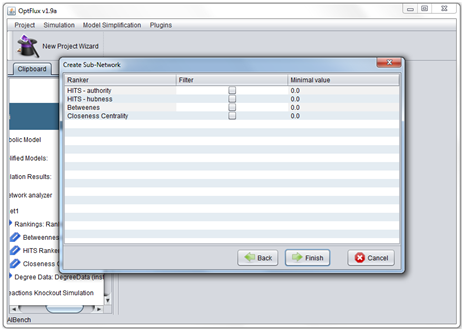
- 4 - The values obtained though any ranking algorithm can be used to define thresholds for vertex removal.
- 5 - Finally vertices and edges can be selected manually for removal.
It should be noted that when a network is filtered a new network is created without the selected elements, but the original network remain unchanged.
Network comparison[edit]
TNA4OptFlux can be used to compare pairs of networks. This functionality was mostly implemented to compare subnetworks derived from the same base though filtering processes. To compare a pair of network go to Plugins →TNA4 → Compare Networks and select the networks. The process of comparison is based in the analysis metrics which were implemented in booth networks and in the vertices which they share. In this situation vertices which have the same id and type are assumed to represent the same biological entity. Besides global network metrics several other parameters are used for the comparison at vertex level:
- 1 - Exclusive vertices and edges: the vertices and edges which appear in only one of the networks being compared are identified.
- 2 - Degree values.
- 3 - Jaccard index .
- 4 - Decision points: these are metabolites which are present in booth networks but that are consumed by different reactions in them. They are called decision points because they can be said to be points where there was an alteration (or decision) of the flux of matter in the system.
- 5 - Average ranking algorithms values, naturally the compared ranking algorithms mast have been applied to the both networks, when necessary these values are normalized.
- 6 - Value of ranking algorithms, normalizes when necessary.
- 7 - Reaction direction: this parameter identifies reversible reactions whose direction differs between networks. It is only of use for networks obtained though simulation filters.
Locating active vertices[edit]
A reaction-compound network in TNA4OptFlux has a structure very similar to a Petri Net. Taking inspiration from this similarity, a functionality called the location of active vertices was included in this tool. It uses an algorithm that starts with a set of “seed” metabolites (the active metabolites set) and from it determines the full set of reactions that can be active when this set of metabolites is present. This functionality works through an iterative process that, at each step, adds the metabolites that can be produced by the reactions in the model by using the current set of active metabolites as substrates to the active metabolites set. Additionally, at each step, all the reactions where all substrates are present in the active metabolites set are added to the active reaction set (which starts empty). The process continues until no further metabolites can be added to the active metabolites set. The final result are sets of reactions which will be active and of metabolites which will exist in the system assuming an unlimited supply of the “seed” metabolites and sufficient time for the reactions to occur. To use this functionality go to Plugins →TNA4 → Locate active vertices, and select a network and “seed” metabolites.
Calculating flux variation[edit]
When studying networks obtained though simulation filtering it can be useful to compare how the flux of the simulations which resulted in the subnetworks differ form on to the other, for this reason TAN4OptFlux has feature that, while not directly related to network analysis, can be of help in the comparison of different simulation results and complement the topological analysis tools. This feature allows a set of simulations to be selected and the results of the comparison among them to be presented both as a table and as bar plots that show the variation of flux values.
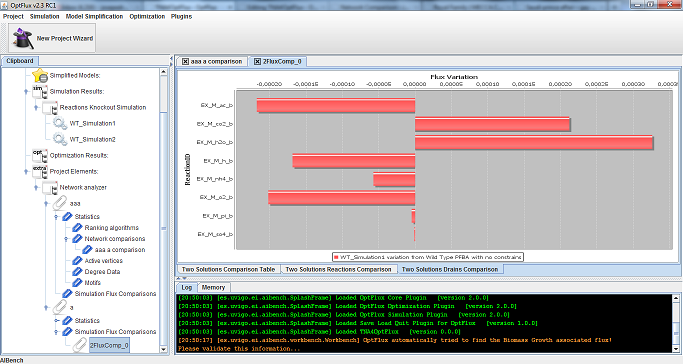
To use this feature go to Plugins →TNA4 → Calculate flux variation, then select a network with a simulation associated (any network which was obtained though simulation filtering) and if it should be compared with a wild type simulation or the simulation associated with another network.
Variation networks[edit]
The concept of variation networks derived from the processes of network comparison and simulation filtering. A variation network is obtained by comparing a pair of networks obtained through simulation filtering, using the same base network and simulation results obtained under different conditions. The result is the creation of a third network containing the elements that differed between the compared networks (i.e. the variations). The idea behind the use of variation networks is to identify the components of a given metabolic system that change when the conditions inherent to the simulations change. Assuming the dimension of these variations to be significantly smaller than the networks themselves, variation networks should be easier to analyze than a full network. The most important decision in this process is to select how the variations between the networks should be identified, i.e. what criteria should be used to decide on the components to be selected. Two methods are included in TNA4OptFlux:
- 1 - Exclusivity: this is the simplest method but it has already proven useful in analyzing phenotype simulation results. The criterion used is based on the identification of the exclusive reaction vertices in each network, i.e. those existing in one of the networks but not the other. After the identification of these reactions, the respective vertices are added to the variation network, together with the vertices corresponding to metabolites that they consume or produce, as well as the edges connecting them.
- 2 - Flux variation: This second method was developed after it was verified that methods based purely on network topology can be, in some cases, insufficient to capture more subtle metabolic flux variations. The flux variation is based in the comparison of flux values of the reactions present in both networks: if the absolute value of the difference in fluxes exceeds a user defined threshold (which can be an absolute flux value or a percentage) then the vertex corresponding to that reaction, as well as the ones corresponding to the metabolites which participate in it and the edges which connect them, are added to the variation network.
These two methods can be used independently or combined. After a variation network is created, it can then be analyzed as any other network using any of the tools available in the plug-in. Besides the comparison of pairs of networks, the concept of variation networks was expanded to work with simulation sets, i.e. sets of phenotype simulations. For creating a variation network from a simulation set, one network not belonging to the set (e.g. obtained from an wild type simulation) is considered the reference network. This network is compared to all networks of the set using the same methods as before. In this case, vertices are only included in the final variation network if they have shown a significant variation in at least K comparisons (where K is a user defined parameter). To obtain a variation network from a pair of network though simulation filtering go to Plugins →TNA4 → Variation Networks → Create varation networks from network pair. Then select the pair of networks to be compared. Finally select the variation criteria, including the flux variation threshold (which can be a relative or absolute value) if this criteria is used. To obtain a variation network from multiple networks first make all necessary simulation and combine them in a set by using Simulation → Create Simulation Set. Then go to Plugins →TNA4 → Variation Networks → Create varation networks from solution set. Select a network obtained though simulation filtering to serve as the reference network and the solution set. Next select the variation criteria and define the K value for each by felling the text box in front of the name of the selected criteria, please note that K must be a value between 0 and 1. Please note that when using a solution set it is only necessary to create the reference network.