Simao.soares (talk | contribs) (→Import genome) |
Simao.soares (talk | contribs) (→Import genome) |
||
| Line 22: | Line 22: | ||
*choose one of the available genomes in The Seed | *choose one of the available genomes in The Seed | ||
*a name for the local annotation | *a name for the local annotation | ||
| − | *the mapping available locally to be used. | + | *the mapping available locally to be used. |
| − | A new genome annotation will be created in the local model-seed with the inserted name. | + | A new genome annotation will be created in the local model-seed with the inserted name.<br /> |
[[File:Import-genome.png]] | [[File:Import-genome.png]] | ||
Revision as of 16:32, 25 March 2013
The Model-seed works in two levels, a local server and The seed server, it is possible to import to the local model-seed:
- biochemistry data
- genome annotations
- mappings
And use these data elements in local operations.
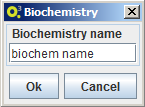
Import biochemistry
To import a biochemistry select the option MSeed -> Import -> Biochemistry and choose a name for the local data element. OptFlux will then ask Model-seed to retrieve the latest biochemistry version from The Seed and to create local of version with the selected name.
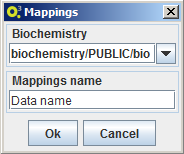
Import mappings
To import a mapping select the option MSeed -> Import -> Mappings and choose a name for the local data element and the local biochemistry to be associated to the new mapping. OptFlux will then ask Model-seed to retrieve the mapping from The Seed and to create a local of version with the inserted name and connected to the selected local biochemistry.
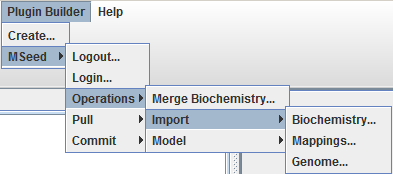
Import genome
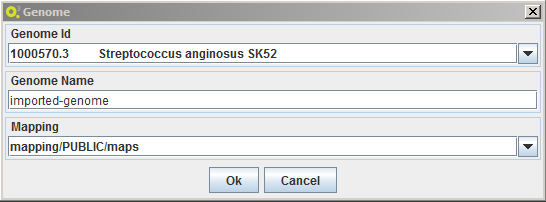
To import a genome select the option MSeed -> Import -> Mappings:
- choose one of the available genomes in The Seed
- a name for the local annotation
- the mapping available locally to be used.
A new genome annotation will be created in the local model-seed with the inserted name.