(→File Definitions) |
(→File Definitions) |
||
| Line 49: | Line 49: | ||
You can also learn more about libSGBN [http://www.sbgn.org/LibSBGN here]. | You can also learn more about libSGBN [http://www.sbgn.org/LibSBGN here]. | ||
<br><br> | <br><br> | ||
| − | <b>KGML</b>: | + | <b>KGML</b>: KGML is an exchange format created for the Kyoto encyclopaedia of genes and genomes ([http://www.genome.jp/kegg/ KEGG]) maps automatic drawing.This format exclusively uses KEGG-ids while genome scale metabolic models do not use these identifiers as primary source of identification of their compounds and reactions, so it is necessary to map the identifiers to make possible the connection between the model and the layout. |
| + | |||
| + | This can be made using the Import KGML interface that the plugin provides: | ||
| + | |||
<br><br> | <br><br> | ||
<b>XGMML</b>: | <b>XGMML</b>: | ||
Revision as of 15:15, 11 November 2013
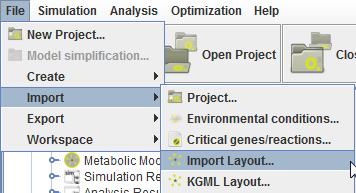
These import layout operations are availabe in the Import menu:
There are 2 major operations for layout importation in OptFlux. The default import operation allows importing layouts from XGMML, COBRA, CellDesigner and SBGN layouts.
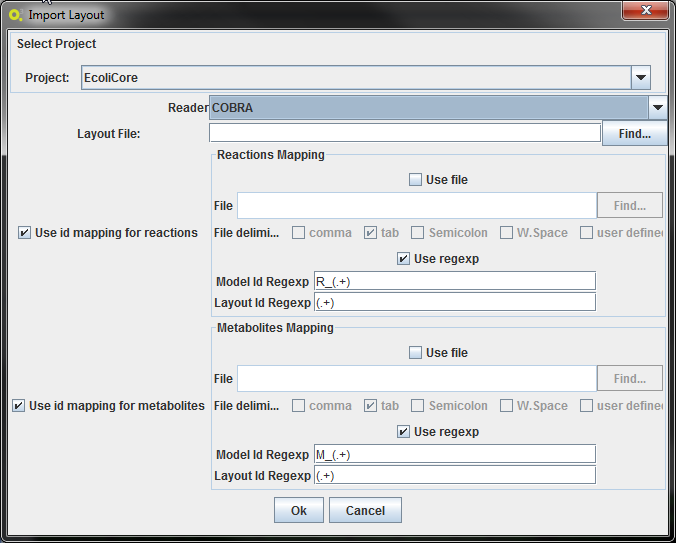
In this interface it is possible to define the mapping of the id's from the metabolic model to the layout. This mapping may not be necessary, but this way it is possible, for instance, to use the same layout for different models.
The mapping can be made two ways:
- Providing a 2-column file: mapping the metabolic model identifier to the layout identifier
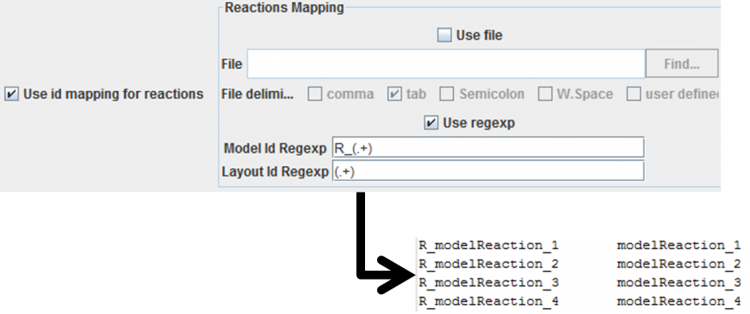
- Applying regexps to the ids: the regexps must contain only one group. If the id's match, by applying those regexp's, then that layout entity will be mapped with the metabolic model entity. An example of a reaction mapping can be seen below.
File Definitions
CellDesigner SBML: CellDesigner (http://www.celldesigner.org/) is a well-known tool for drawing cellular networks, based on the process diagram paradigm, with a graphical notation system proposed by Kitano. These are stored using an extension of the Systems Biology Markup Language (SBML).
COBRA Maps:
SBGN PD: SBGN (http://www.sbgn.org/) is a visual language developed by biochemists, modellers and computer scientists that aims at being the standard for visual representation of biological processes.
SBGN defines three visual languages: Process Description (PD), Entity Relationship (ER) and Activity Flow (AF). This plugin supports PD language files. You can learn more about these maps by downloading this file.
At the moment SBGN-PD maps are only supported entities of the type "simple chemical" (for metabolites) and "process" (for reactions). Clone nodes are considered currency metabolites.
You can also learn more about libSGBN here.
KGML: KGML is an exchange format created for the Kyoto encyclopaedia of genes and genomes (KEGG) maps automatic drawing.This format exclusively uses KEGG-ids while genome scale metabolic models do not use these identifiers as primary source of identification of their compounds and reactions, so it is necessary to map the identifiers to make possible the connection between the model and the layout.
This can be made using the Import KGML interface that the plugin provides:
XGMML: